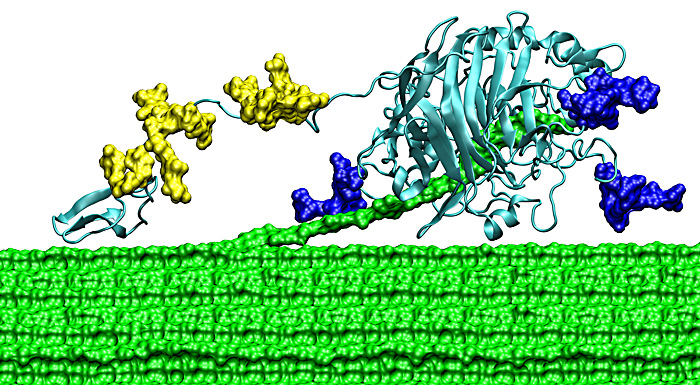
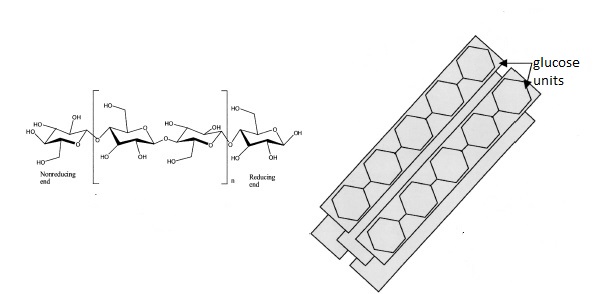
In the last post (yes, its been a while!) I talked about free cellulases. The problem with free cellulases is recovery and that you need several different types of cellulases to work together to accomplish the breakdown of cellulose. What if you could have a scaffold that did that for you? Bacteria and fungi have done just that and its called the cellulosome! Of course, its a complex structure but the basics are illustrated in Figure 1.
 |
| Figure 1 (taken from ref 1) |
|
|
The cellulosome consists of several different parts. First a protein known as scaffoldin is attached to the bacterial (or fungal) cell. Scaffolding contains domains known as cohesins and carbohydrate binding proteins (binding to cellulose for example) and a domain with surface layer homology of unknown function (2004); cohesins bind a second type of protein called dockerins and dockerins bind the cellulosome enzymes that degrade carbohydrates, cellulose being just one of those carbohydrates. There are different types of cohesins (type I-III in 2004) and they bind different classes of dockerins (types I-III) through a small binding site of four amino acids. Which dockerin binds to which cohesin is highly specific. Figure 1 shows a fairly simple representation of a cellulosome. Other micro-organisms have more complex structures involving 3 different scaffoldin proteins that link together.
Interestingly, the presence of cellulosome genes in an organisms genonome does not necessarily mean it can degrade cellulose. Developing an efficient and robust cellulosome containing the enzymes that work effectively together is then the job of the lignocellulose biochemist. To this end, researchers have been working on designing minicellulosomes.
We already know that Saccharomyces cerevisea is a organism of choice in the fermentation of glucose to produce ethanol due to its tolerance for high ethanol concentration (see blog post#). S. cereviseae is also a good choice because its genetics are well studied with many tools available for gene manipulation. Further, yeast is able to "display" proteins on its surface. i.e. it can display a cellulosome on on its surface meaning that not only will it do the expression but it puts the whole system together and presents it on its surface. The scientist, doesn't need to purify the proteins and put them together herself. Because of the genetic tractability, different cohesins from different organisms can be mixed and matched to produce the most efficient cellulosome, unavailable naturally. Tsai et al, 2009, put together cohesins from 3 different bacterial strains and displayed them on the yeast cell surface (figure 2). The antibodies (upside down green and blue and brown Y shapes with yellow or green stars) were used to detect the expression of the different components ofn teh surface of the yeast.
 |
Figure 2. Functional assembly of minicellulosomes on
the yeast cell surface. A trifunctional scaffoldin (Scaf-ctf)
consisting of an internal CBD flanked by three divergent cohesin (C)
domains from C. thermocellum (t),
C. cellulolyticum (c),
and R. flavefaciens (f)
was displayed on the yeast cell surface. Three different cellulases (E1,
E2, and E3) fused with the corresponding dockerin domain (either Dt, Dc,
or Df)
were expressed in E. coli. Cell lysates containing these cellulases were
mixed with yeast cells displaying Scaf-ctf for the functional assembly of the
minicellulosome.
|
The authors showed that this cellulosome was 2.6x more efficient than if the enzymes were added in their soluble form and ethanol production was 95% of its theoretical value!
References:
1.
Roy H. Doi1
&
Akihiko Kosugi. 2004 Cellulosomes: plant-cell-wall-degrading enzyme complexes Nature Reviews Microbiology 2,
541-551 (July 2004)